Howard Chang (Stanford, HHMI) 2: LncRNA Function at the RNA Level: Xist
Or Chang I'm, a professor at Stanford University. And investigator, with the Howard Hughes, Medical Institute. This. Is the second part of my talk and, I'll be focusing on long non-coding RNAs. The. New century brought a major mystery, to biologists. With. A completion, the human genome project while. Just got busy studying, the different kinds of RNAs, that get made or transcribed, from the genome and there. Are classes, of RNA like this one where we take the sequence run. Into the computer, who can immediately see, that this, is a protein coding gene they're. Well-known domains, and, we immediately infer. Specific. Functions, for. The particular protein product, but. There are also other, transcripts. Very, abundant. And very numerous that. You. Know look quite, the same coming, off the sequencer, but, when you run into the computer, you get a big question mark and these. Are not to be long, non-coding RNA, genes these. Transcripts, were also, made by RNA, polymerase 2 they're, spliced, sometimes, Polident, elated yet. Their functions, were quite mysterious and so, lesson questions about could they be impossible, in both and human. Diseases, or other important, traits and finally. Could there be a systematic. Code do, you help us understand, this new class of sequences. This. Particular challenge is, really like trying to decode, a lost language and. We can learn from history about, how that's done. This. Is the rosetta stone a very important, archaeological. Artifact, that, was uncovered by Napoleon's, soldiers from. In Egypt, and. The. Language, shown, here are additional, hieroglyphics. Common. Gyptian in Greek. Egyptian. Hieroglyphics, were basically, a lost language, we, can see them on ancient monuments, but people couldn't read them and this, stone, was very important, because at the same text, 3, times once. Each of these three languages and by. Going back and forth between these three versions. People. Can see that, there are certain characters, that, was show up when, certain words are mentioned in the other Lane which for example the. Name Ptolemy, was show up with, the following hieroglyphics. And by, the systematic, correspondence. People. Could work out actually. A code of transformation. From, hieroglyphics, to, characters. That we actually understand, so. Not only can we read the text on the rosetta stone we, can actually read all the text in ancient monuments, written, hieroglyphics, and this. Is a very apt analogy, because, RNA, like. These characters, fold, up into complicated, shapes so. Our task is really to assign form, to, function understand. There's systematic relationships. This. Particular feel is really undergoing, a revolution because. Of the rapid, pace of discovery of. RNA, genes we. Now know that human genome encodes, nearly. 60-thousand. Long, non-coding RNA. Genes and I used, abbreviation, linked RNA for, long non-coding RNAs, and this. Includes very classic, examples, of RNA. Start discover in the 90s - more recent, examples, some, of them are shown here as, the. Typical. Example, so the first no, examples, acted, on chromatin, that's by no means their only function.
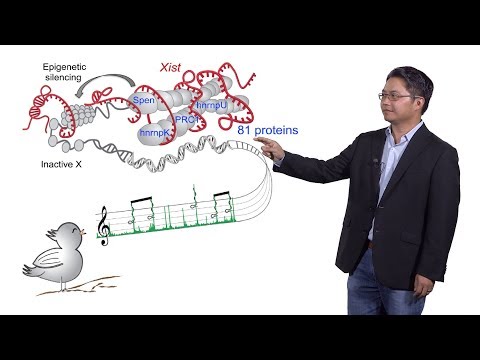
And. The approach that my, lab has taken has focused, on delving, into mechanisms. Of a select, number of long non-coding RNAs, and also developing technologies, to help, other investigators. Tackle, their, favorite linked RNA genes and, today. I want to talk to you about a very classic. Linked RNA and that is exists the, master, regulator of X chromosome inactivation. Let. Me remind you that. Men. And women are different and men, have, an X and a Y chromosome at least mammals. And females. Have, two X chromosomes. So. To make the gene expression from this. Second. X equivalent. To, this tiny Y in. Mammals. One. Of the X chromosomes, needs to be shut down and that is, done by this long non-coding RNA, gene exists, is. Transcribed. Only, about one of the x chromosome, it's, 17, kilobases, law and it spreads and coats the, inactive, X chromosome, and somehow, induces. Epigenetic. Silencing on, that chromosome so, there's much less gene, transcription, on the inactive, X there's. A small number of genes that escape X inactivation and, so, these escapees, define, sex differences, between, men and women, in biology and, disease. This. Image on the right here shows in situ hybridization, of exists. You can see that's very unusual because it's, strictly, in the nucleus and I only just coats that one chromosome that creates a classic, cytological, structure called the bar body. Each. Cell, in, the female body makes, it a choice a random, choice about which X chromosome, wants to shut down and, that has very interesting consequences. For, example in this calico, cat these, different patches of color. And on the fur our consequence. Of this random X inactivation, that's. Happening, in each, individual. Skin cells which, then grow up to make these little patches, of clones. So. It was long believed that exists. Would act with protein partners, but. These partners have remained elusive for a long time and. We. Approach this question by, developing, an RNA, directed proteomic. Approach would, you call chirp, MS, chirp, mass spectrometry, the, idea is to basically, fix. The, RNA protein complex, in the living cell and then, retrieve, the specific, RNA, with, complimentary, óscar nucleotides, we're gonna subject the associated. Proteins, to, mass spectrometry. This. Reveal that, exists. As 17, kilobase. Long RNA is associated, with 81, proteins. Additional. Work suggested, that ten of these interactions, proteins. Are probably direct, RNA. Protein interactions, the. Remaining, interaction, of probably through indirect, protein. Protein interactions. And. So this, set of proteins really gave a parts, list for all the different jobs that exist has to do including, spreading. Across the X chromosome, causing. Gene silencing in changing, the morphology of the chromis. How. Do we figure out which, are the key proteins, out of this big list of 81. Proteins, here. Were were aided by some classical, genetics, were. From Anton woods and colleague has shown that, a small, sequence, on exists, called a repeat, i was, necessary, for gene silencing so. On the 17, kb long RNA these, few hundred bases actually, were absolutely, required for gene silencing in, avery. People ishe mutant could. Still be made still, spread across the chromosome, encoded but, it wouldn't silence the genes so. We knew that something very important, was probably, on the a repeat. So, we perform, chirp. Mass spectrometry. On either wild-type cells or, on the a repeat, deletion, mutant the. Bottom graph shows a scatter, plot of the peptide. Counts, in our results, in the full length on the, x-axis, or the avery people lesion you can, see that everything, was on the 45, degree diagonal exactly, the same except, for three proteins, whose peptide, count fall to zero in. The a repeat deletion, so, this show really that exists, RNA it was a modular. Ly organized, different. Piece of the RNA had different partners and these, three proteins. Likely. And needed to a repeat, for interaction.
So. Now that we've sort of narrowed, down the list, of interesting candidates, we, can further go on to show that by individually. Knocking, down these, candidate, factors that this protein called span was, very, important, for gene silencing on X chromosome, and subsume. Were from several other groups independently, confirm, this result. Okay. Spen, was a factor, that was initially, described. From, Drosophila, genetics, it's. Involved, in development, and it, had never been implicating X chromosome inactivation before. Or linked RNA function, but actually it was the perfect candidate, its. Protein, structure, is very suggestive on the, n-terminus, there, are four professional, binding, RNA, binding RM, domains. And the, c-terminus, there's, a spock t'me that, binds to chromatin silencing factors, the. Plant homolog. Spen, is, involved in silencing, transposon. Their parasitic. DNA elements and. So this is an ancient protein. That's, been commandeered, to, do this job and mammalian or eutherian. Specific, job in X chromosome inactivation. The. Humans, spend protein, it actually is it associates. With a repressor, chromatin, repressor complex, called, the nerd complex. And together, with enzymes like histone, deacetylase, three methyl. And binding. Domains involved, in DNA methylation and. So these are factors are associating gene silencing and. I'll refer the audience, to the talk by David Allis in, eye biology, on histone, deacetylases. About. Chromatin. In gene silencing and finally. We actually subsequently. Show that, the RM domains of spen interact. Directly with the a repeat. Okay. So. To. Summarize a fairly, large body of work we discover that the set of proteins, that interact would, exist, ashlee, assemble. In parts, X. Chromosome, inactivation in. Exists ashin happens, when embryonic stem cells differentiate, so. Before, that differentiation, happens, in pluripotent, stem cells one, set of factors associate. With exists already with. Differentiation, a second, set of factors including spen. Associate. With exist and that, completes. The reconstitution, of the exist protein, complex. And. We believe that there is a logic, to this division of labor of these proteins, this, first set of proteins that assembles. Including, the polycomb, complex they're. Involved, in epigenetic. Memory they're maintaining, the status quo so, if you're an active element you'll stay active your fear silent, you'll stay silent, so, what is needed to turn a previously, active, x chromosome, into the inactive, x is the involvement of factors like spent, and associated.
Proteins They, will deactivate. An. Active, elements. And once, they're deactivated. You can push those elements, into, this maintenance module to, remember, that state and perpetuate. The gene silencing. There. Like histone, marks associated which are these steps which. Provides. A different readout, of their activity. It. Turn out that RNA, structure, is critical, for understanding how, exists, interacts, with each of its partners. RNA. Is a single-stranded. Flux mamala we can base pair with itself and other molecules in the, fundamental. Unit of RNA structure, is that duplex, two, strands of RNA base, pairing together in an Watson Crick base pairing, and, interaction, and so. We developed a method to track, these RNA duplexes, in living, cells using. A chemical called psoralen and this, method is called sorrel, analysis, of RNA interaction, of structure or Paris, for short in this. Method we. Introduce, psoralen, which diffuse into cells and cross, link the two strands together in the living cell, we. Can then isolate these. Purities. Like joint, fragments. Ligate. The two ends of the RNA together by. Proximity ligation once. We remove the psoralen. When, we the, two ends are ligated, we can sequence, them and, when we sequence, every. Read gives. A single, molecule evidence, that these two strands were touching each other in the living cell and in, this this interaction, is shown like this kind of Arc you see at the bottom ok so the more arcs means there are more interactions. Using. This methodology, we, could interrogate, the, structure. That exists, so shown, at the bottom is sort of the human exist locus little bit longer 19. KB and there's, a series of repeats, which will describe, in the, primary, sequence of, the exist molecule, but, on the top is the secondary, structure map and these are being the RNA duplexes, and what we discovered, is first of all that there are many long-range, interactions, that spent over a spear, occurring, over kilobases. The. Interactions. Are modular, you see loops within loops so their domains, of RNA structure that are revealed and, finally, we see that there are certain regions that can adopt, several, different, structures, they're, called alternate structures we see one. Base. Interacting with this and other base that, can't happen at the same time so these are alternate, structures, okay. This. Provide this proved to be quite important, for understanding, it's. An RNA light exists for, example the a repeat, the. A repeat, derives as named because, a the, same sequence, repeats itself, at eight and a half times in. That. Exist, sequence and so, there have been a number of computational. And biochemical, approaches, trying, to look at the a repeat, structure, but, because you have these repeating. Identical, units these. One-dimensional, methods. Will, give sort, of multiple, equivalent. Solutions, and therefore. We need to a two dimensional. Method like Paris to help us resolve that, kind of paradox, and so, here's what the pair of structure showed and each, arc again is one of these duplex, maps that were detected, in the living cell and. I show really that the a repeat. Region. Folds. Up as follows, basically. The unit, of interaction, is the it's.
Basically The inter, it's between two repeats. It's. The inter repeat duplex that is staggered, okay so repeat, one interacts. With the repeat two three. Reaches, over to five and four, over to eight and that corresponds, to this big arc over here and then, six, again to seven. When. Spend binds the, interaction. Surface, the cross-linking size, exactly, of a junction, of the two duplexes, between. The single and the double-stranded regions, and here, is this and therefore of these and. Here's what a single unit will look like the red color is a single, copy of the of the a repeal, one here and in, a second one again here. Okay and the, asterik mark is the is. The interaction, sighs so the staggered, duplex is the unit of interaction. Okay. Let's. Turn now to look at the chromatin, consequences. Of this, link RNA action, so. When we think about protein, coding genes of disease genes we think about the coding sequence, but for each gene, there, are DNA elements switches. That decide when and where that gene turns, on and off and these, elements, are the, binding sites of, protein-based. Transcription. Factors, and of regulatory RNAs. In. The human genome the pattern actually looks more like this just, two percent of the real stay is protein coding and the vast majority of, the, intergenic. Region, is actually this non-coding, space with potentially. Many DNA regulatory, elements, we, also know that most. Of the disease, variants. Associated with human disease occur. In this non-coding, space. To. Interrogate, the non-coding genome, it's. Important, to remember that, in. Every human cell two, meters of DNA is packed into a 10 micron nucleus. Mostly. DNA's highly. Wound up not accessible. Not. Available, to the cells machinery only, the active elements, can be read and simply, find this accessible, elements, give us a lot of information about. The. Gene regulatory program, in the cell, several. Years ago my colleague, will Greenleaf and I as Stanford developed, this methodology, called, a taxi, assay. Of transposes, accessible, chromatin, it, uses an enzyme, which. Copies, and pastes DNA and we've, loaded into this enzyme, already DNA. Sequences, that go onto our sequencing, machine, when. This enzyme, reacts. With the periodic chromatin, it, can only copy and paste into the open chromatin sites but not the compact, its elements, and therefore. In a single, step you selectively, in covalently, tag, they're accessible, elements, of, interest you can then amplify, in sequence. And. This method led to a pretty, amazing a million fold improvement in the sensitivity, and a hundredfold. Improvement, in the speed of mapping. Regulatory, landscape, in. In, biology. For. The interested. Audience, I refer, to the part 1 of my talk on epigenomic. Technologies, will Reidel more into this particular, technology. But. In the context, of exists. We. Can use this technology to ask how, does exist silence, one chromosome, but not the other in the same nucleus, to. Do that we have to use some genetic tricks to follow. Cells, from. A. An animal model where. The, chromosome, from mom in, a chromosome from dad have, sequence, differences, so, they're so-called hybrid animal. We, can do an allele, specific mapping, once, we see accessible, elements, we can also look at those sequence, differences, to ask is it coming from the mom chromosome, or the dad chromosome, and this. Showed us that on the. X chromosome which is shown from, left to right here, okay. The add the active X has, lots of accessible, elements, and, these included genes and their associated regulatory. Elements but, the inactive, x has largely, lost, this, accessibility. Except. For. The genes that escaped X inactivation okay. That includes exists itself, and a few other genes are highlighted, here and we further discovered, that when genes escaped exact, X inactivation it's. Not the entire gene in its regulatory unit, it's just, the promoter, but not the long-range distal, enhancers, that escaped, as an activation so the promoters actually the the unit that decides how, escape happens, what makes men, and women potentially, different. Okay. Now, we always have the ambition of not, only just knowing, which elements, are being controlled, active, or not active accessible. Or not accessible but. To know their spatial, organization within. The three-dimensional context of the cell and this involves a further innovation, in that technology which.
We Call a taxi. So. I told you that attacks seek is basically, spray-painting. The genome with DNA, adapters, what. We decided we could do is that we can actually add a little floor floor basically is a fluorescent, molecule at the end of the DNA adapter, now, once we spray-paint, the cell we, can actually image, the. Pattern of accessible elements in situ, and then, afterwards. Sequence. Figure. Out what the elements that we've been imaging so, a taxi. Basically. Is a very easy way to remember like you're seeing with your eyes okay. So. The next image is an example of an attack C of. A cancer, cell the red colors attack see the accessible, elements they're blue colors that be all the DNA, she, want you to notice that the, the. The accessible, sized person just fill up the entire nucleus, is not homogeneous they're, actually, regions, are more accessible than, other ones that are not in a dark in, the red channel I'll show you that there's a very specific organization. So. Now we can revisit the, question of X inactivation with, this lens with this viewpoint and. On. The bottoms what I've shown you before already, with attack seek active. X chromosome, loss of accessible, sites inactive. X loss. Of accessible sites but, in our taxi, now in the green color we, can see that in fact the enacted ax is actually focused, the red color is the exist RNA, okay so you see that they exist everywhere they exist RNA is there. Is a black, hole of accessibility. I'm. Gonna Tuggle. To, this other view so you can see that basically, if you go back and forth got, really every place is red color is the RNA. Is gone okay. And furthermore, that. The RNA. And the black claws moved to the edge of the nucleus, which they where the inactive X chromosome, typically, resides, so. This tells us that in fact, exists. RNA has local has organized. A local, three-dimensional, chromatin, structure and basically, pulled in active acts into, that sort. Of nuclear location. Finally. We can gain some insight, into the evolutionary, origin. The potential origin, of this power for long non-coding RNA. I told. You about span the critical, partner for gene silencing for, exists and, when we discovered, is that when, we remove spent.
Knock It out and now, look at autosomes, all the other chromosomes, besides, X there. Are actually, hundreds, of DNA elements that, become accessible, become, active, and nearly. Half, of these elements are curricula. Class, of, endogenous. Retroviruses. Call, ERV K these. Are ancient, parasites. Viruses that. Infected. Invading, into mammalian genomes and then spread, and copy themselves and we discovered, that spend is actually, part of this virus fighting. Machinery, it. Recognizes, an RNA from the virus and. Basically trust me sighs, shut it down by, epigenetically. Silencing. It by chromatin, modification. This. Is a very interesting finding, because this. Little sequence, from, the Irv Kay that, attracts, spent turns. Out to be quite similar, to the sequence, of the Avery P that I've been telling you about it. Was recognized, in 2008. That. Exists. Evolved, from a series of transposon. Irv integrating. Into the locus and then, the sequence of the a rupees shown in red is quite, similar to the the herbs that are being targeted by spent on autosomes, this. Is led, us to a a model, then that, in fact. Spend. RNA interactions, are ancient and, sort. Of conserved feature and when, an ancient, virus, jump. Into exists. On the X chromosome a proto exists, this. Endow. That sequence, with the capacity, to attract spend a powerful. Epigenetic, machinery, when, this sequence the subsequent, amplify, and mobilize, it, repurposed. This link RNA, into. This powerful machine for, silencing, an entire chromosome, and so therefore. Those leads to hypothesis. Of X inactivation by, viral, mimicry, in effect, the female, cell treats. The inactive, X like, a big heap, of viruses, the. Female cells pretends, that there's a raging var infection, going, on on the inactive X chromosome. You can then recruit, this very powerful machinery. To, shut that down chromosome, to accomplish the goal of dosage, compensation. Therefore. In summary I've told you about long. Non-coding rnas, and chromatin. And this. Sketched by MC, Escher showing. The two hands drawing each other is a very apt, analogy because, we think that RNA. And chromatin, this, these two polymers, basically. Template, each other the, information is, copy from one backward, to the other and furthermore. Like, these two hands, we like to think that the linked RNA could be a driver, of evolutionary. Innovation. In gene regulation. You.
2020-01-30 00:30


